Transcription: Start and Stop
In this lesson, we learn about transcription, when and where does transcription start, when and where does transcription stop, and how does transcription work.
DNA Transcription and The Central Dogma
What is the “central dogma”? The central dogma of molecular biology is basically the idea explaining how proteins are made, from transcription of the DNA to the translation of mRNA. The central dogma includes the following essential processes: DNA transcription, DNA replication, mRNA translation, reverse transcription, and protein synthesis.
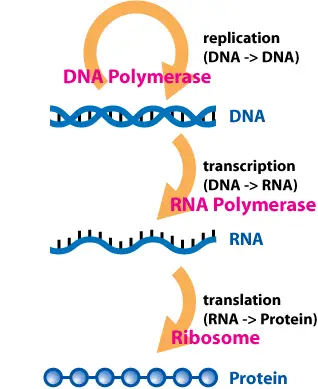
3 Major Steps in DNA Transcription (RNA synthesis)
Transcription, also known as RNA synthesis, is the process of making the mRNA from the DNA. There are 3 stages to DNA Transcription: 1) initiation, 2) elongation, and 3) termination.
- Step 1: Initiation. At the promoter region, the enzyme RNA polymerase unwinds the DNA at its promoter region. This creates an opening for the RNA polymerase to initiate the beginning of transcription.
- Step 2: Elongation. During elongation, the RNA polymerase runs down the template strand and copies the DNA by matching the DNA’s nucleotides with its complementary. Essentially, the addition of nucleotides to the growing mRNA strand is the step elongation.
- Step 3: Termination. As the RNA polymerase goes down the template strand, the unwounded DNA rewinds into its original configuration. Transcription stops at the termination site, which is the last step of transcription, termination.
There are two strands of our DNA: the coding strand and the template strand.
The coding strand runs from 5′ to 3′. The template strand is its opposite, its antiparallel; it runs from 3′ to 5′.
Remember these nucelotide complementaries:
-
A ——- T (DNA)
-
A ——-U (RNA)
-
C——–G (DNA)
The RNA polymerase makes the mRNA strand that runs from 5′ to 3′.
- Coding Strand: 5′ to 3′
- Template Strand: 3′ to 5′
- mRNA Strand (RNA polymerase runs in this direction): 5′ to 3′
Here is an example of what each strand would look like during the transcription process.
DNA Template: [[ 3‘ ATTCCGGAA 5‘ ]]
mRNA Strand: [[ 5‘ UAAGGCCUU3‘]]
DNA Coding: [[ 5‘ TAAGGCCTT 3‘ ]]
If you’re taking a test and it asks you to find the codes for the mRNA strand, a fast way to figure out the mRNA strand’s letters is to just look at the coding strand and switch all of the T’s for U’s. Don’t forget to label its head 5′ and bottom end 3′!
Start and Stop of Transcription:
- Start: Promoter Region
- Stop: Termination Region (usually a row of T’s)
- Why is the termination region of transcription usually a row of T’s? Great question! The complementary nucleotide of T is A, and there are always two hydrogen bonds between the T and A. Unlike the nucleotides T and A, complementary C and G are bonded to each other with three hydrogen bonds. Logically, two hydrogen bonds are weaker than three hydrogen bonds. Thus, it is easier for the RNA polymerase to fall off of the termination region and be released from the DNA.
- Eukaryotes vs. Prokaryotes? For eukaryotes, termination factors are required in order for the mRNA to be released, but for prokaryotes, termination factors are usually not required. The termination factors are molecules that bind to the termination signal region and causes the RNA polymerase to fall off. Again, the RNA polymerase begins to fall off along a string of approximately 300 base pairs, which are usually nucleotides of T.
- As for prokaryotes, the mRNA strand has an inverted repeat along with a string of Us . Like T’s, U’s form only double bonds with A’s, and it is likely that the RNA polymerase can fall off more easily along this row of weaker bonds. Additionally, the inverted repeat in the mRNA folds into a loop structure, further disrupting the binding of the RNA polymerase to the DNA at hand. In general, there are 2 types of termination for prokaryotes and bacteria. Read more about Intrinsic Terminators and Rho-independent transcription termination here.
Transcription could also synthesize other types of RNAs. Besides mRNA, transcription is also the production of miRNA, tRNA, and rRNA.
Exons and Introns
Many eukaryotes have exons and introns within their mRNA strands right after transcription.
Exons are also known as coding regions, and introns are known as noncoding regions. Thus, introns are the ones removed from the mRNA because they are parts that do not code for anything.
- Exons: Coding regions —-> not removed
- Introns: Noncoding regions —-> removed
Eukaryotes vs Bacteria Prokaryotes: Start and Finish of Transcription
Where does transcription start and end? How does a RNA polymerase know where to attach to the DNA and start transcribing?
RNA polymerases recognize transcriptions sites slightly differently between bacteria and eukaryotes. For prokaryotes, RNA polymerase searches the DNA template strand for promoters. Promoters are sites on DNA that tell the polymerase where to bind and start transcription. For eukaryotes, RNA polymerases need the help of transcription factors to bind to the promoter. Transcription factors first bind, and then RNA polymerase is initiated to bind to the spot as well. This forms the transcription pre-initiation complex (PIC).
[Read more about differences between Prokaryote and Eukaryote DNA Transcription here.]
Bonus Simulation Time! 🙂
Input your own DNA sequence into our Auto-Transcription Simulation, and sit back and relax! DNA sequences are automatically converted into mRNA sequences. This project was made by our students using Scratch programming. For a tutorial on instructions, click here. To play the simulation, click the thumbnail picture below.
Copyright 2018 Moosmosis: All Rights Reserved
Please Like our Facebook page to support our open-access youth education initiatives! 🙂
Source:
- “Transcribe and Translate a Gene.” Learn Genetics. Web. <http://learn.genetics.utah.edu/content/molecules/transcribe/>.
- “Transcription and Translation.” Transcription and Translation. Web. 28 Mar. 2016. <https://www.genome.gov/27552603>.
Happy Learning! 🙂
Categories: Biology










![Central Chemoreceptor vs Peripheral Chemoreceptor in Respiration [Biology, MCAT, USMLE]](https://moosmosis.files.wordpress.com/2022/02/respiratory-control_med.jpeg?w=200&h=200&crop=1)


Thank you for posting. I will share this with my students.
LikeLiked by 1 person
Thank you so much for sharing, Bob! We hope your students like our post too! Have a wonderful New Year’s, to you and your wonderful students. Cheers!
LikeLike
Nice overview! I’m a molecular biologist and found it to be accurate 🙂 Very useful to see a summary of the differences between prokaryotes and eukaryotes.
LikeLiked by 1 person
Thank you so much, jkay! How awesome to hear from a molecular biologist! 🙂 We strive to be as accurate as possible, with peer-review edits before publishing, and we’re always open to feedback for improvement. You’ve got an amazing sustainable environmental project going, and just wanted to let you know that your Green Stars Project on ethical consumerism is inspiring to us students. Happy New Years!
LikeLike
Interesting post. Thanks for stopping by my blog and following. 🙂 — Suzanne
LikeLiked by 1 person
Thank you for stopping by as well, Susan! Your experiences and stories in India and beyond are fascinating. Looking forward to more!
LikeLiked by 1 person