Prokaryote DNA Transcription and Bacteria Transcription
Prokaryote Transcription Steps
There are three major steps to DNA transcription in prokaryotes and bacteria. The three steps are: 1. Initiation, 2. Elongation, and 3. Termination.
- Step 1 Bacteria Transcription: Sigma Factor scans for initiation site or promoter sequence. Opens complex, unwinds DNA to start
- Step 2 Bacteria Transcription: RNA polymerase selects NTP complementary to DNA template, catalyzes phosphodiester bonds. Performs elongation in transcription bubbles.
- Step 3 Bacteria Transcription: RNA polymerase stops at termination signals (See below for the two types of termination signals for prokaryotes and bacteria)
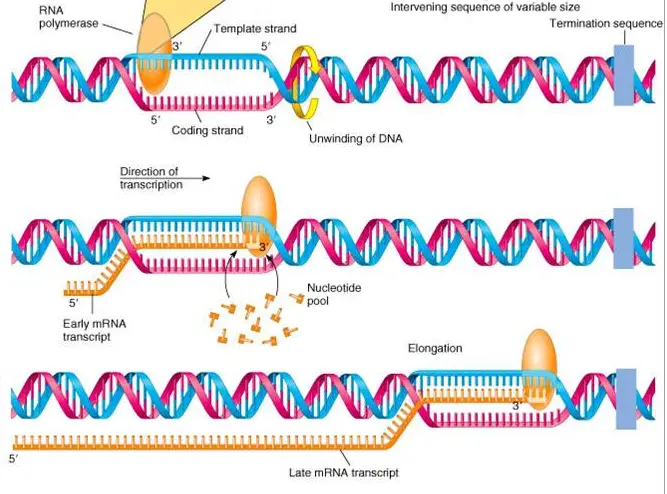
Prokaryote Transcription vs. Eukaryote Transcription: What are the differences?
1 Major Difference: Transcription in bacteria is less complex than transcription in eukaryotes.
Bacteria have promoter regions, a region on the DNA that tells RNA polymerases where to start. RNA polymerases are weakly attached to the DNA and then zip off down the template strand to transcribe. The double helix is opened up immediately, exposing the DNA’s nucleotides and allowing the RNA polymerase to then complement nucleotides accordingly.
RNA polymerases have the ability to transcribe left-right or right-left, but must travel in the direction of 5′ to 3′ (which means that the RNA polymerase travels down a DNA template strand of 3′ to 5′).
RNA polymerase’s mRNA strand [[5‘ to 3‘]]
DNA Template Strand [[3‘ to 5‘]]
DNA has two strands, so which strand is the template strand and which one is the coding strand?
The answer actually depends on which strand the RNA polymerase attaches onto and the direction of the bacterial sigma factor.
2nd Major difference between prokaryotic transcription and eukaryotic transcription: The sigma factor only exists in bacteria, not eukaryotes.
The sigma factor is a subunit of bacterial polymerase; eukaryotes do not have sigma factors during their transcription process.
Placed in front of the RNA polymerase, the bacterial sigma factor recognizes the promoter region and actually starts transcribing 10 nucleotides. After transcribing 10 nucleotides, the sigma factor latches off and allows the RNA polymerase to move forward to transcribe. The RNA polymerase continues elongation, which catalyzes phosphodiester bonds and elongation in transcription bubbles.
The RNA polymerase continues to transcribe until it reaches a termination signal or sequence, usually the termination sequence is a row of T’s or A’s. To understand why the termination sequence is a row of T’s or A’s, we have to look at their molecular bonds. Nucleotide T and A create the weakest hydrogen bonds, only two hydrogen bonds between each other, compared to C and G, that have three hydrogen bonds. Thus, it is easier to break off from these weaker bonds, and this concept serves ideal for the end of transcription.
As transcription ends, the bacterial sigma factor returns and reattaches to the freed RNA polymerase. Both the RNA polymerase and sigma factor go search for a new promoter to start the transcription process again.
What are the 2 types of termination for DNA Transcription for Prokaryotes and Bacteria? How do bacteria terminate DNA Transcription?
Prokaryotes and bacteria terminate DNA transcription and RNA synthesis via 2 types: 1. Intrinsic terminators, and 2. Rho-dependent terminators.
- Intrinsic Terminators: These are sequences that cause DNA transcription to terminate without any other factors. The DNA contain palindromes or stretches of T’s or A’s in DNA template. The resulting mRNA thus has a UUUUU-like stretch. Uniquely, the mRNA also has a GC rich loop that folds on itself before this stretch of UUUU to the DNA’s usual stretch of T’s. This GC rich loop is a RNA hairpin, telling the RNA polymerase to pause and destabilize the RNA-DNA hybrid and terminate DNA transcription.
- Rho-dependent Terminators: These DNA transcription terminator types need the rho factor (helicase) to terminate DNA transcription. The rho protein binds to a C rich, G-poor RNA chain and pulls the RNA polymerase away from the DNA template. ATP is needed for the Rho protein to terminate DNA transcription for these bacteria and prokaryote types.
Read more on Eukaryotic DNA Transcription in our Next free lesson here.
Copyright 2018 Moosmosis: All Rights Reserved
Please Like our Facebook page to support our open-access youth education initiatives! 🙂












Really helpful info on transcription!! Thanks
LikeLiked by 1 person
You’re very welcome, Sam! We’re glad to help.
LikeLike